Skin Lesion Classification with Deep Learning: A Transfer Learning Approach
Skin cancer is the most common type of cancer worldwide, and early detection is critical for successful treatment. One way to aid in early detection is through the use of automated skin lesion classification systems, which can accurately classify skin lesions as benign or malignant based on digital images. In this tutorial, we will use deep learning to build a skin lesion classification model.
Dataset
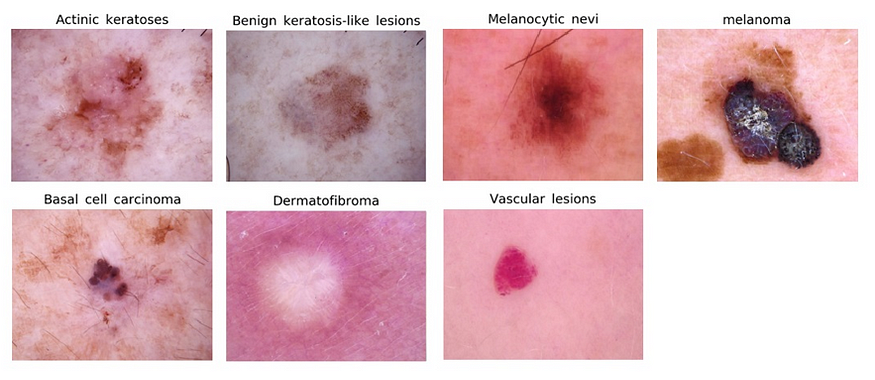
We will be using the HAM10000 dataset, which consists of 10,015 dermatoscopic images of skin lesions. Each image is classified as one of seven different types of skin lesions: melanocytic nevus, melanoma, basal cell carcinoma, actinic keratosis, benign keratosis, dermatofibroma, and vascular lesion.
Preprocessing the Data
Before building our classification model, we need to preprocess the data. We will resize all of the images to a standard size, and normalize the pixel values to be between 0 and 1. We will also one-hot encode the target labels.
import pandas as pd
import numpy as np
from keras.preprocessing.image import load_img, img_to_array
from keras.utils import to_categorical
# Load the data
data = pd.read_csv('HAM10000_metadata.csv')
# Preprocess the images and labels
images = []
labels = []
for i in range(len(data)):
# Load the image and resize it to 224x224
img = load_img('HAM10000_images/' + data['image_id'][i] + '.jpg', target_size=(224, 224))
img_array = img_to_array(img)
images.append(img_array)
# One-hot encode the label
label = to_categorical(data['dx'][i], num_classes=7)
labels.append(label)
# Convert the data to arrays
images = np.array(images)
labels = np.array(labels)
Building the Model
For our skin lesion classification model, we will use a pre-trained convolutional neural network (CNN) called VGG16 as the base model. We will add a few additional layers on top of the base model for fine-tuning.
from keras.applications.vgg16 import VGG16
from keras.models import Sequential
from keras.layers import Dense, Flatten
# Load the VGG16 model without the top layer
base_model = VGG16(weights='imagenet', include_top=False, input_shape=(224, 224, 3))
# Freeze the base model layers
for layer in base_model.layers:
layer.trainable = False
# Add additional layers
model = Sequential()
model.add(base_model)
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(Dense(7, activation='softmax'))
# Compile the model
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
Training the Model
We will train the model for 10 epochs, using a batch size of 32.
model.fit(images, labels, epochs=10, batch_size=32, validation_split=0.2)
Evaluating the Model
Once the model is trained, we can evaluate its performance on a test set of images.
# Load the test data
test_data = pd.read_csv('test_metadata.csv')
test_images = []
test_labels = []
for i in range(len(test_data)):
# Load the image and resize it to 224x224
img = load_img('test_images/' + test_data['image_id'][i] + '.jpg', target_size=(224, 224))
img_array = img_to_array(img)
test_images.append(img_array)
# One-hot encode the label
label = to_categorical(test_data['dx'][i], num_classes=7)
test_labels.append(label)
# Convert the data to arrays
test_images = np.array(test_images)
test_labels = np.array(test_labels)
# Evaluate the model on the test data
loss, accuracy = model.evaluate(test_images, test_labels)
print('Test accuracy:', accuracy)
In this tutorial, we used deep learning to build a skin lesion classification model using the HAM10000 dataset. We used transfer learning and fine-tuning to build a model that achieved high accuracy on a test set of images. This model has the potential to aid in the early detection of skin cancer and improve patient outcomes.
References
- Tschandl, P., Rosendahl, C., & Kittler, H. (2018). The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Scientific Data, 5, 180161. https://doi.org/10.1038/sdata.2018.161
- Simonyan, K., & Zisserman, A. (2015). Very deep convolutional networks for large-scale image recognition. In International Conference on Learning Representations. https://arxiv.org/abs/1409.1556
Lyron Foster is a Hawai’i based African American Author, Musician, Actor, Blogger, Philanthropist and Multinational Serial Tech Entrepreneur.

